Abstract
Introduction:
Proteins, despite being the primary effectors of cellular processes, are often studied only indirectly through analysis of the transcriptome. However, it is clear that the relationship between mRNA expression and protein expression is approximate at best. In Acute Myeloid Leukemia (AML), the genome and transcriptome have been thoroughly characterized, but the proteome has been less well studied. Here, we present a deep-scale study of the proteomes of 44 primary AML bone marrow samples representing a wide range of AML across the spectrum of cytogenetic risk, common mutations, and driver fusions.
Methods:
Bone marrow samples were collected at presentation from 44 adult patients with de novo AML as part of an institutional banking protocol, and buffy coat cells were immediately cryopreserved without further manipulation. Cryovials were thawed in the presence of the cell permeable serine protease inhibitor diisopropyl fluorophosphate (DFP) to inactivate the abundant neutrophil serine proteases (ELANE, CTSG, PRTN3, and PRSS57), and further processed for nano-liquid chromatography mass spectrometry in the presence of an extensive cocktail of protease inhibitors. Both label-free quantification (LFQ) and tandem-mass-tag (TMT) deep-scale proteomics were performed on these 44 patient samples, as well as 3 lineage-depleted bone marrow samples from healthy adult donors. Matching RNA-seq and exome sequencing data were available for the same samples as part of The Cancer Genome Atlas (TCGA) AML project.
Results:
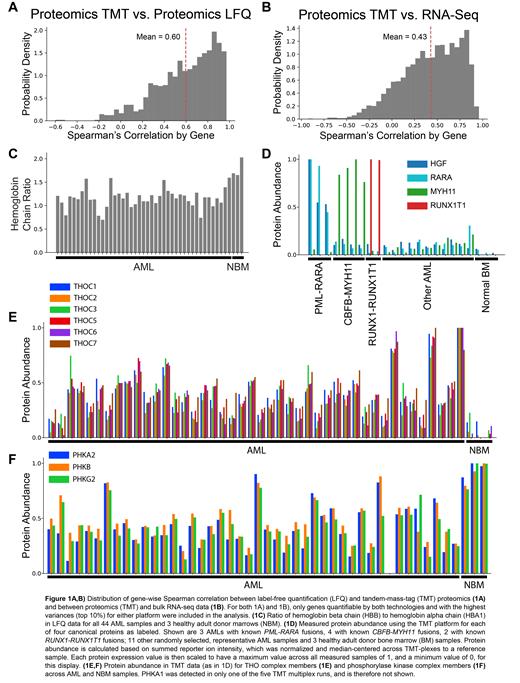
10,651 and 6,679 unique proteins were detected in the TMT and LFQ experiments, respectively. Correlations between measurements derived from the independent proteomic platforms (i.e. TMT and LFQ) is higher (mean Spearman correlation, 0.60, Figure 1A) than correlation between proteomic (TMT) and transcriptomic measurements from bulk RNA-seq data (Spearman 0.43, Figure 1B). Quality checks of the proteomic data strongly supported the reliability of quantification of protein measurements; for example, the mean ratio of beta globin protein (HBB) to alpha globin (HBA1) was 1.2 +/- 0.25 (Figure 1C), and several proteins known to be dysregulated by specific AML-initiating fusion proteins (for PML-RARA, HGF and RARA; for RUNX1-RUNX1T1, RUNX1T1; and for CBFB-MYH11, MYH11) were detected in the expected samples (Figure 1D). Globally, 1,364 proteins were differentially expressed in the AML samples (corrected p-value <0.05, fold change ≥ 1.5) compared to the lineage-depleted, healthy bone marrow samples. Globally overexpressed proteins were enriched for ribosomal RNA modification, mitochondrial protein import, nuclear export, and the mitochondrial electron transport chain, among others. These overexpressed proteins include 61 cell surface proteins that could potentially represent therapeutic targets (overexpressed on average in 82% of AML samples, range 25-97%). Globally downregulated proteins in AML samples were enriched for glycogen metabolism and protein groups associated with mature neutrophils (reflecting the expected maturation block in AML), among others. 771 of the 1364 differentially expressed proteins (56.5%) showed only minimal variability in mRNA expression levels (fold change of <1.1 between AML and normal marrow CD34 cell mRNA) that could not explain dysregulated protein expression. Several protein complexes likewise showed coordinated differential expression in the proteomic data, but no change in the transcriptome, including the THO complex (Figure 1E) and the phosphorylase kinase complex (Figure 1F), among others, indicating the presence of posttranscriptional regulation of the levels of many proteins in AML samples.
Conclusion:
We have created a deep-scale proteomic database from a set of well-characterized AML samples, allowing for a proteogenomic study of AML. We have identified many examples of post-transcriptional regulation of key metabolic pathways that may be relevant for better understanding AML cell metabolism and therapeutic vulnerabilities. Additional studies linking patterns of protein dysregulation with a variety of AML covariates are underway.
No relevant conflicts of interest to declare.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal